Fleming Fund Regional Grant SeqAfrica
Extending whole genome capacity for AMR surveillance
Project Overview (2019–2026)
SeqAfrica was a UK Fleming Fund regional grant project aimed at strengthening Whole Genome Sequencing (WGS) and bioinformatics capacity across Africa to enhance antimicrobial resistance (AMR) surveillance. Led by the Technical University of Denmark (DTU), with key partners including the National Institute for Communicable Diseases (NICD) in South Africa, the Noguchi Memorial Institute for Medical Research (NMIMR) in Ghana, and the Institut Pasteur de Dakar (IPD) in Senegal, the project supported three regional sequencing centers. These centers processed bacterial isolates from across the continent, enabling outbreak investigations and tracking of resistance genes across human, animal, and environmental sectors under a One Health approach. SeqAfrica also provided technical assistance, training, and simulated exercises. The project aimed to reduce reliance on external sequencing services, build local expertise, and foster sustainable genomic surveillance infrastructure in Africa.
During Phase I (2019-2023), SeqAfrica established the regional sequencing network with regional WGS RCs based in Ghana, Nigeria, Tanzania, and South Africa, building on already established WGS and bioinformatics capacity and capabilities. Throughout Phase I, the project worked towards further developing, expanding, and supporting WGS and bioinformatics capacity for AMR surveillance across Africa using short-read technology, at the regional WGS RCs and beyond. In Phase II (2023 – 2026), the project aimed to maintain core services with a downsized network and a moderate sequencing capacity for AMR surveillance with an increased focus on virtual training, data use, and dissemination. The project also piloted a few sentinel sites that focused on long-read sequencing using MinION sequencers from ONT paired with an offline laptop-based analysis designed for non-technical users, offering a proof of concept for democratizing WGS
Section: Accomplishments
-
Regional genomic surveillance network
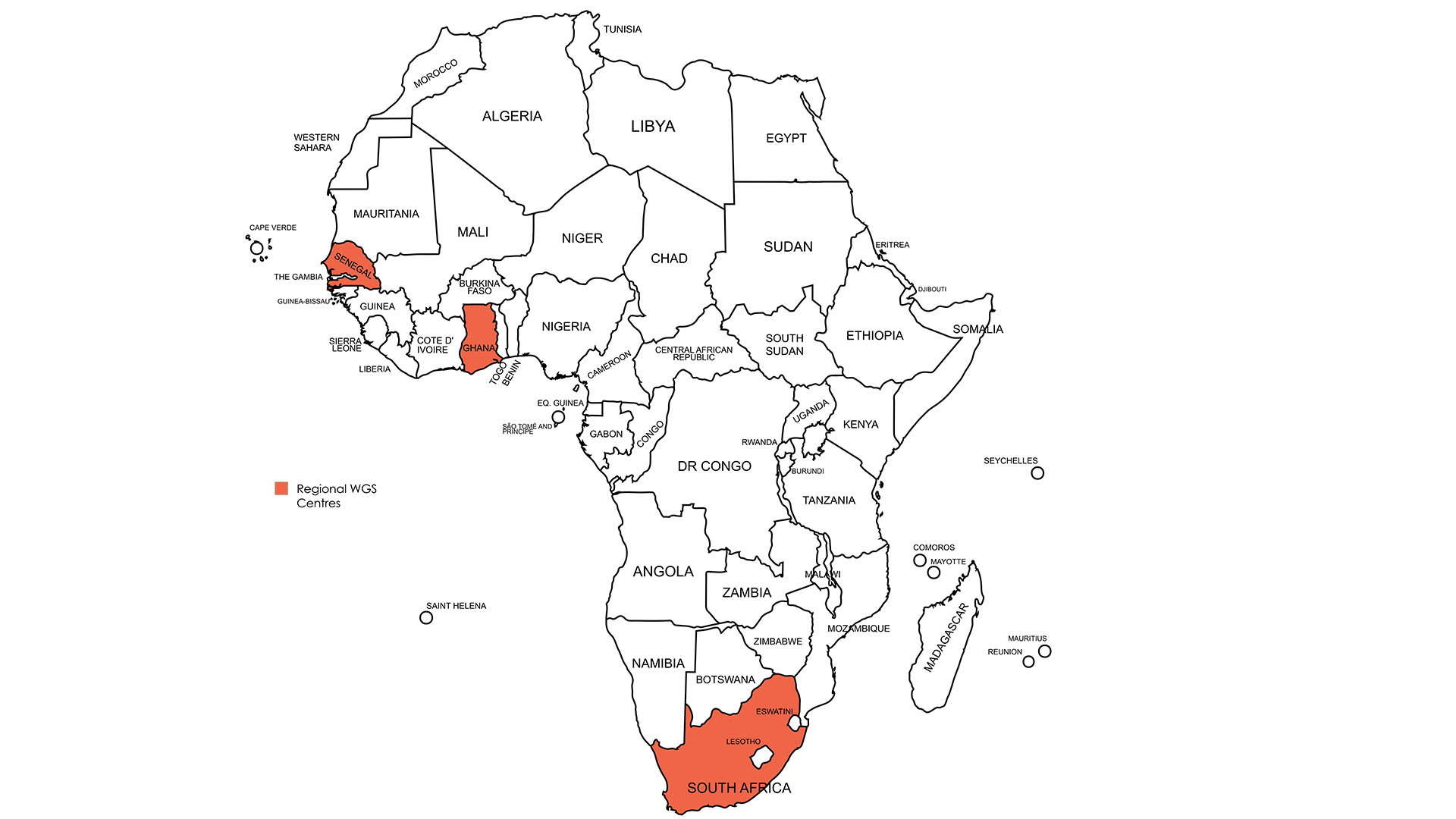
Established three regional sequencing hubs in South Africa (NICD), Ghana (NMIMR), and Senegal (IPD), supported by portable Oxford Nanopore (ONT) field sequencing sites to expand access across settings. -
Large-scale genomic sequencing impact
Generated over 36,000 genomes, including 25,000+ bacterial genomes and 10,000+ SARS-CoV-2 genomes across 24 countries, supporting 60+ publications and informing surveillance, research, and policy. -
Capacity building & training
Trained 200+ researchers and laboratory staff through workshops, simulation exercises, and bioinformatics training, strengthening long-term regional expertise in pathogen genomics. -
Community of Practice
Maintained a monthly community of practice (Apr 2024–Mar 2026) with international experts. The network will transition into the African Bioinformatics Institute’s Pathogen Genomics and AMR Community of Practice. -
Expanded Salmonella surveillance
In South Africa, Salmonella genomes increased from 847 (2020) to 12,459, placing the country among the global leaders in publicly available Salmonella genomic data. -
Rapid response during COVID-19
Sequencing infrastructure rapidly pivoted to SARS-CoV-2 surveillance, enabling near real-time variant tracking. NICD played a key regional role in supporting outbreak monitoring and public health response.
